SCI Publications
2025
C. Scully-Allison, K. Menear, K. Potter, A. McNutt, K.E. Isaacs, D. Duplyakin.
“Same Data, Different Audiences: Using Personas to Scope a Supercomputing Job Queue Visualization,” 2025.
Domain-specific visualizations sometimes focus on narrow, albeit important, tasks for one group of users. This focus limits the utility of a visualization to other groups working with the same data. While tasks elicited from other groups can present a design pitfall if not disambiguated, they also present a design opportunity -- development of visualizations that support multiple groups. This development choice presents a trade off of broadening the scope but limiting support for the more narrow tasks of any one group, which in some cases can enhance the overall utility of the visualization. We investigate this scenario through a design study where we develop \textitGuidepost, a notebook-embedded visualization of supercomputer queue data that helps scientists assess supercomputer queue wait times, machine learning researchers understand prediction accuracy, and system maintainers analyze usage trends. We adapt the use of personas for visualization design from existing literature in the HCI and software engineering domains and apply them in categorizing tasks based on their uniqueness across the stakeholder personas. Under this model, tasks shared between all groups should be supported by interactive visualizations and tasks unique to each group can be deferred to scripting with notebook-embedded visualization design. We evaluate our visualization with nine expert analysts organized into two groups: a "research analyst" group that uses supercomputer queue data in their research (representing the Machine Learning researchers and Jobs Data Analyst personas) and a "supercomputer user" group that uses this data conditionally (representing the HPC User persona). We find that our visualization serves our three stakeholder groups by enabling users to successfully execute shared tasks with point-and-click interaction while facilitating case-specific programmatic analysis workflows.
S. Sebbio, L. Carnevale, D. Balouek, M. Parashar, M. Villari.
“Data-Driven Operational Artificial Intelligence for Computing Continuum: A Natural Disaster Management Use Case,” In 2025 IEEE 25th International Symposium on Cluster, Cloud and Internet Computing Workshops (CCGridW), pp. 92-99. 2025.
The increasing frequency of documented natural disasters can be attributed to advances in communication technologies, such as satellites, the Internet, and smart devices that facilitate better disaster reporting. This is coupled with an actual rise in the occurrence of such events and improved documentation of their impacts. These trends underscore the pressing need for scalable and intelligent technological solutions to efficiently process large datasets, allowing informed decision-making and effective disaster response. This study presents a Computing Continuum framework that integrates intelligence across cloud, edge and deep edge tiers for efficient disaster data processing. A significant characteristic is the incorporation of Artificial Intelligence for IT Operations (AIOps), which leverages machine learning and analytics to facilitate dynamic resource management and adaptive system modeling, thereby addressing the intricate challenges posed by disaster scenarios. The architecture encompasses an AI-driven framework for monitoring and managing service, network, and infrastructure layers, tailoring policies to specific disaster needs. The proposed framework is applied to wildfire management, leveraging an AI Operation Manager to coordinate sensor-equipped drones for real-time data acquisition and processing. Operating at the deep edge tier, these drones transmit environmental data to edge and cloud infrastructures for analysis. This multi-tiered approach improves situational awareness, disaster response, and resource utilization.
M. Shao, S. Joshi.
“Domain-Shift Immunity in Deep Deformable Registration via Local Feature Representations,” Subtitled “arXiv:2512.23142v1,” 2025.
Deep learning has advanced deformable image registration, surpassing traditional optimization-based methods in both accuracy and efficiency. However, learning-based models are widely believed to be sensitive to domain shift, with robustness typically pursued through large and diverse training datasets, without explaining the underlying mechanisms. In this work, we show that domain-shift immunity is an inherent property of deep deformable registration models, arising from their reliance on local feature representations rather than global appearance for deformation estimation. To isolate and validate this mechanism, we introduce UniReg, a universal registration framework that decouples feature extraction from deformation estimation using fixed, pre-trained feature extractors and a UNet-based deformation network. Despite training on a single dataset, UniReg exhibits robust cross-domain and multi-modal performance comparable to optimization-based methods. Our analysis further reveals that failures of conventional CNN-based models under modality shift originate from dataset-induced biases in early convolutional layers. These findings identify local feature consistency as the key driver of robustness in learning-based deformable registration and motivate backbone designs that preserve domain-invariant local features.
H. Shrestha, J. Wilburn, B. Bollen, A.M. McNutt, A. Lex, L. Harrison.
“ReVISitPy: Python Bindings for the reVISit Study Framework,” In Eurovis 2025, 2025.
User experiments are an important part of visualization research, yet they remain costly, time-consuming to create, and difficult to prototype and pilot. The process of prototyping a study-from initial design to data collection and analysis-often requires the use of multiple systems (e.g. webservers and databases), adding complexity. We present reVISitPy, a Python library that enables visualization researchers to design, pilot deployments, and analyze pilot data entirely within a Jupyter notebook. Re- VISitPy provides a higher-level Python interface for the reVISit Domain-Specific Language (DSL) and study framework, which traditionally relies on manually authoring complex JSON configuration files. As study configurations grow larger, editing raw JSON becomes increasingly tedious and error-prone. By streamlining the configuration, testing, and preliminary analysis workflows, reVISitPy reduces the overhead of study prototyping and helps researchers quickly iterate on study designs before full deployment through the reVISit framework.
X. Tang, X. Li, T. Tasdizen.
“Dynamic Scale for Transformer,” In Medical Imaging with Deep Learning, 2025.
To enhance the hierarchical transformer with fixed embedding sizes, we propose a dynamic CLS token that aggregates information from CLS tokens across all layers, each embedded with varying receptive fields, by leveraging a squeeze-and-excitation module. This architecture offers a more flexible approach to utilizing multi-scale features in transformers. Code is available at: https://github.com/xiaoyatang/DynamicCLS.git.
X. Tang, B. Zhang, M.M. Ho, B.S. Knudsen, T. Tasdizen.
“DuoFormer: Leveraging Hierarchical Representations by Local and Global Attention Vision Transformer,” Subtitled “arXiv:2506.12982,” 2025.
Despite the widespread adoption of transformers in medical applications, the exploration of multi-scale learning through transformers remains limited, while hierarchical representations are considered advantageous for computer-aided medical diagnosis. We propose a novel hierarchical transformer model that adeptly integrates the feature extraction capabilities of Convolutional Neural Networks (CNNs) with the advanced representational potential of Vision Transformers (ViTs). Addressing the lack of inductive biases and dependence on extensive training datasets in ViTs, our model employs a CNN backbone to generate hierarchical visual representations. These representations are adapted for transformer input through an innovative patch tokenization process, preserving the inherited multi-scale inductive biases. We also introduce a scale-wise attention mechanism that directly captures intra-scale and inter-scale associations. This mechanism complements patch-wise attention by enhancing spatial understanding and preserving global perception, which we refer to as local and global attention, respectively. Our model significantly outperforms baseline models in terms of classification accuracy, demonstrating its efficiency in bridging the gap between Convolutional Neural Networks (CNNs) and Vision Transformers (ViTs). The components are designed as plug-and-play for different CNN architectures and can be adapted for multiple applications. The code is available at https://github.com/xiaoyatang/DuoFormer.git.
L.C.R. Tanner, A. Busatto, J.A. Bergquist, W.W. Good, B. Zenger, G. Plank, A. Narayan, K. Gillette, R.S. MacLeod.
“Uncertainty quantification via polynomial chaos expansion of myocardial fibre orientation and cardiac activation patterns,” In The Journal of Physiology, Wiley, 2025.
Predictive models and computational simulations of cardiac electrophysiology depend on precise anatomical representations, including the local myocardial fibre structure. However, obtaining patient-specific fibre information is challenging. In addition, the influence of physiological variability in fibre orientation on cardiac activation simulations is poorly understood. We implemented rule-based algorithms to generate fibres and robust uncertainty quantification methods to determine model output variability with respect to ventricular activation sequences. We used polynomial chaos, which reduces computational demands by using an emulator to approximate the underlying forward model. Our study examined activation sequences in response to nine stimuli and five metrics quantifying essential features of the activation sequence. The results indicated that the primary fibre orientation impacts the overall spread of activation, which could impact more complex patterns of activation; however, there is minimal impact on the location of discrete activation features, such as breakthrough sites. For free wall stimuli, the standard deviation (STD) was highest near the stimulus site, diminishing with distance. Apical stimuli showed complementary STD patterns, with epicardial pacing maximizing STD in the right basal area and endocardial pacing in the left. Ventricular junction stimuli exhibited symmetrical STD patterns, low near the stimulus but increasing sharply towards the apex, peaking on the left in the apical region. Furthermore, variability in the imbrication or helix angle did not impact the activation sequences. We conclude that in many relevant modelling contexts, the variability in myocardial fibre orientation can play an important role in the resulting activation sequences and should be accounted for.
L.C.R. Tanner, A. Busatto, T. Grandits, J.A. Bergquist, B. Zenger, S. Pezzuto, G. Plank, R.S. MacLeod, K. Gillette.
“Reconstructing ventricular activation sequences from epicardial data: Insights from Geodesic Back-Propagation optimization in porcine models,” In Computers in Biology and Medicine, Vol. 198, 2025.
Cardiac digital twins (CDTs) are emerging as powerful tools in personalized medicine, providing subject-specific models to simulate and understand cardiac function. A central challenge in constructing CDTs is accurately personalizing the structure and function of the His-Purkinje system (HPS), which determines ventricular activation. In this study, we leveraged a novel modified Geodesic-BP method to infer early activation sites (EASs) from epicardial activation times. The EASs can then serve as a surrogate for Purkinje-myocardial junctions, facilitating anterograde ventricular activation. We used both experimental porcine (N 5) and synthetic (N 5) datasets of epicardial activation times measured or assigned to locations on an electrode sock. For both datasets, we optimized for initial estimates of 5, 50, 100, and 200 EASs and assessed output variability by repeating the inference process 10 times. Assessments were based on matching the predicted activation times at both the epicardial locations and from intracardiac measurements made with multielectrode needles, and throughout the ventricular myocardium for the synthetic dataset. The algorithm could consistently recover global ventricular activation patterns from epicardial data alone. For the experimental dataset, the minimum and maximum mean absolute differences were 0.19 ms and 3.86 ms on the epicardial sock and 2.65 ms and 10.69 ms for the needles. For the synthetic dataset, the corresponding values were 0.13 ms and 2.81 ms on the sock and 2.42 ms and 14.07 ms ms throughout the ventricular myocardium. However, discrepancies between the epicardial surface and intramural myocardium, overfitting of EASs, and variability across repeated runs revealed key limitations. These findings highlight both the overall potential and current limitations of inferring EASs using the proposed optimization approach. They demonstrate the feasibility of deriving informative activation patterns from limited data, while underscoring the need to incorporate stronger physiological priors and anatomical constraints. Ultimately, our results motivate future efforts to refine simulation-based personalization frameworks, improve robustness, and enhance the physiological realism of CDTs for more accurate and reliable applications.
W. Tao, S. Joshi, R. Whitaker.
“Integrated Model Selection and Scalability in Functional Data Analysis through Bayesian Learning,” Subtitled “Preprints.org,” 2025.
DOI: 10.20944/preprints202503.0658.v1
Functional data, including one-dimensional curves and higher-dimensional surfaces, have become increasingly prominent across scientific disciplines. They offer a continuous perspective that captures subtle dynamics and richer structures compared to discrete representations, thereby preserving essential information and facilitating more natural modeling of real-world phenomena, especially in sparse or irregularly sampled settings. A key challenge lies in identifying low-dimensional representations and estimating covariance structures that capture population statistics effectively. We propose a novel Bayesian framework with a nonparametric kernel expansion and a sparse prior, enabling direct modeling of measured data and avoiding the artificial biases from regridding. Our method, Bayesian scalable functional data analysis (BSFDA), automatically selects both subspace dimensionalities and basis functions, reducing computational overhead through an efficient variational optimization strategy. We further propose a faster approximate variant that maintains comparable accuracy but accelerates computations significantly on large-scale datasets. Extensive simulation studies demonstrate that our framework outperforms conventional techniques in covariance estimation and dimensionality selection, showing resilience to high dimensionality and irregular sampling. The proposed methodology proves effective for multidimensional functional data and showcases practical applicability in biomedical and meteorological datasets. Overall, BSFDA offers an adaptive, continuous, and scalable solution for modern functional data analysis across diverse scientific domains.
W. Tao, T. J. Somorin, J. Kueper, A. Dixon, N. Kass, N. Khan, K. Iyer, J. Wagoner, A. Rogers, R. Whitaker, S. Elhabian, J. A. Goldstein.
“Quantifying Sagittal Craniosynostosis Severity: A Machine Learning Approach With CranioRate,” In The Cleft Palate Craniofacial Journal, Sage, 2025.
DOI: 10.1177/105566562513473
Objective
Design
Setting
Participants
Interventions
Main Outcomes
Results
Conclusions
Y. Tatari, H.T. Nguyen, A. Arzani, P. Newell .
“Investigation of particle transport in geothermal systems using integrated CFD–DEM and data-driven approaches,” In Geothermics, Vol. 136, Elsevier, 2025.
Geothermal systems provide continuous, low-carbon energy by harnessing the Earth’s renewable heat, but their efficiency can be hindered by issues such as limited heat production and thermal breakthrough. A promising approach to overcome these issues is to inject polymer-based microcapsules into fractures to modify permeability, which requires a clear understanding of particle transport within the fracture network. To do so, this study used Computational Fluid Dynamics–Discrete Element Method (CFD–DEM) simulations combined with machine learning (ML) to capture particle transport behavior under varying temperature conditions. The dataset comprises 45 CFD–DEM test cases, which are generated by coupling OpenFOAM (for fluid dynamics) and LIGGGHTS (for particle tracking), enabling detailed modeling of thermo-hydro processes and particle interactions. To assess their influence on transport behavior, key parameters include particle diameter (), formation temperature (), and particle volume fraction (). Supervised learning models, including random forest and interpretable decision tree classifiers, were trained to classify flow blockage. Feature importance analysis identified and as the most critical factors impacting the particle transports. To avoid sealing progression, the accumulation of low-velocity particles over time was fit with a sigmoid function. Results show that higher particle concentrations and larger diameters reduce transport efficiency, while elevated inlet velocities enhance particle mobility and prolong transport through the fracture. This interpretable data-driven approach, grounded in CFDEM simulations, offers a predictive tool for particle transport in fractures subject to complex geothermal environments.
M. Taufer, R. Mihalcea, M. Turk, D. Lopresti, A. Wierman, K. Butler, S. Koenig, D. Danks, W. Gropp, M. Parashar, Y. Gil, B. Regli, R. Rajaraman, D. Jensen, N. Bliss, M. Maher.
“Now More Than Ever, Foundational AI Research and Infrastructure Depends on the Federal Government,” Subtitled “arXiv:2506.14679,” 2025.
Leadership in the field of AI is vital for our nation's economy and security. Maintaining this leadership requires investments by the federal government. The federal investment in foundation AI research is essential for U.S. leadership in the field. Providing accessible AI infrastructure will benefit everyone. Now is the time to increase the federal support, which will be complementary to, and help drive, the nation's high-tech industry investments.
T. Transue, B. Wang.
“Learning Decentralized Swarms Using Rotation Equivariant Graph Neural Networks,” Subtitled “arXiv:2502.17612,” 2025.
The orchestration of agents to optimize a collective objective without centralized control is challenging yet crucial for applications such as controlling autonomous fleets, and surveillance and reconnaissance using sensor networks. Decentralized controller design has been inspired by self-organization found in nature, with a prominent source of inspiration being flocking; however, decentralized controllers struggle to maintain flock cohesion. The graph neural network (GNN) architecture has emerged as an indispensable machine learning tool for developing decentralized controllers capable of maintaining flock cohesion, but they fail to exploit the symmetries present in flocking dynamics, hindering their generalizability. We enforce rotation equivariance and translation invariance symmetries in decentralized flocking GNN controllers and achieve comparable flocking control with 70% less training data and 75% fewer trainable weights than existing GNN controllers without these symmetries enforced. We also show that our symmetry-aware controller generalizes better than existing GNN controllers. Code and animations are available at github.com/Utah-Math-Data-Science/Equivariant-Decentralized-Controllers.
J.P. Tuffour, R. Ewing, G. Tian.
“Augmenting Low-Carbon Sustainable Cities Using Transit-Linked Mixed-Use Designs: A Quasi-Experimental Regression of Travel and Emission Outcomes,” In Sustainable Cities and Society, Vol. 135, Elsevier, 2025.
Cities today continue to struggle with the converging twin crises of rising vehicular air pollution and emission-intensive land use, prompting a renewed urgency for climate-smart urban designs such as mixed-use (MXDs) and transit-oriented developments (TODs). Yet, not all developments are designed to have a transit-focused orientation, and even if they are, individual lifestyle choices confound their true effects. This study investigates whether those MXDs situated near high-frequency transit (a design typology we call Transit-Oriented Mixed-Use Developments (TOD-MXDs)) represent a more sustainable antidote to the environmental and mobility crises afflicting cities. Leveraging the largest geo-spatially harmonized database of individual-level and built environment characteristics across 36 diverse U.S. regions, we employ a mix of quasi-experimental design and advanced multivariate regression modeling to estimate travel characteristics and emissions outcomes. Results showed that although TOD-MXDs remain relatively scarce, they consistently outperform their non-transit-oriented counterparts, generating about 20 % less VMT, higher walking (87.6 %) and biking (83 %) trip shares, and significantly lower CO2e emissions, even after accounting for residential self-selection and the full suite of 7-D built environment variables. Furthermore, a unit increase in the presence of transit around mixed-use designs was associated with a ∼13.8 % reduction in VMT, suggesting that coupling land-use diversity with transit integration produces a synergistic effect that advances low-carbon and sustainable city outcomes. As conventional planning and policymakers strive for climate-resilient and healthy communities, this study makes a bold case: TOD-MXDs are not just better, they may be essential to the future of climate adaptation in cities through built-environment design.
J. Turnage, M. Lowery, J. Jakeman, Z. Morrow, A. Narayan.
“An Optimal Weighted Least-Squares Method for Operator Learning,” Subtitled “arXiv:2512.11168v1,” 2025.
We consider the problem of learning an unknown, possibly nonlinear operator between separable Hilbert spaces from supervised data. Inputs are drawn from a prescribed probability measure on the input space, and outputs are (possibly noisy) evaluations of the target operator. We regard admissible operators as square-integrable maps with respect to a fixed approximation measure, and we measure reconstruction error in the corresponding Bochner norm. For a finite-dimensional approximation space  of dimension
of dimension 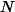 , we study weighted least squares estimators in
, we study weighted least squares estimators in  and establish probabilistic stability and accuracy bounds in the Bochner norm. We show that there exist sampling measures and weights - defined via an operator-level Christoffel function - that yield uniformly well-conditioned Gram matrices and near-optimal sample complexity, with a number of training samples
and establish probabilistic stability and accuracy bounds in the Bochner norm. We show that there exist sampling measures and weights - defined via an operator-level Christoffel function - that yield uniformly well-conditioned Gram matrices and near-optimal sample complexity, with a number of training samples  on the order of
on the order of 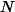
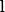


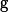
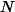 . We complement the analysis by constructing explicit operator approximation spaces in cases of interest: rank-one linear operators that are dense in the class of bounded linear operators, and rank-one polynomial operators that are dense in the Bochner space under mild assumptions on the approximation measure. For both families we describe implementable procedures for sampling from the associated optimal measures. Finally, we demonstrate the effectiveness of this framework on several benchmark problems, including learning solution operators for the Poisson equation, viscous Burgers' equation, and the incompressible Navier-Stokes equations.
. We complement the analysis by constructing explicit operator approximation spaces in cases of interest: rank-one linear operators that are dense in the class of bounded linear operators, and rank-one polynomial operators that are dense in the Bochner space under mild assumptions on the approximation measure. For both families we describe implementable procedures for sampling from the associated optimal measures. Finally, we demonstrate the effectiveness of this framework on several benchmark problems, including learning solution operators for the Poisson equation, viscous Burgers' equation, and the incompressible Navier-Stokes equations.
M. Usman, P.N. Castillo, A. Narayan, L.H. Timmins.
“Integrating Uncertainty Quantification into Computational Fluid Dynamics Models of Coronary Arteries Under Steady Flow,” Subtitled “arXiv:2512.11017,” 2025.
Computational models are continuously integrated in the clinical space, where they support clinicians in disease diagnosis, prognosis, and prevention strategies. While assisting in clinical space, these computational models frequently use deterministic approaches, where the inherent (aleatoric) variability of input parameters is ignored. This questions the credibility and often hinders the clinical adoption of these computational models. Therefore, in this study, we introduced uncertainty quantification in the computational fluid dynamics models of the left main coronary artery to analyze the influence of input hemodynamics parameters on wall shear stress (WSS). UncertainSCI was used, where an emulator was built using polynomial chaos expansion between the input parameters and the output quantity of interest, and the output sensitivities and statistics were directly extracted from the emulator. The uncertainty-informed framework was first applied to an analytical solution of the Navier-Stokes equation (Poiseuille flow) and then to a patient-specific model of the left main coronary artery. Different input hemodynamics parameters are considered, such as pressure, viscosity, density, velocity, and radius, whereas wall shear stress was considered as our output quantity of interest. The results suggest that velocity dominated the variability in WSS in the analytical model (~79%), whereas viscosity dominated in the patient-specific model (~59%). The results further suggest that out of all the Sobol indices interactions, unary interactions were the most dominant ones, contributing ~93.2% and ~99% for the analytical and patient-specific model, respectively. This study will enhance confidence in computational models, facilitating their adoption in the clinical space to improve decision-making for coronary artery disease diagnosis, prognosis, and therapeutic strategies.
A.L. Veroneze Solórzano, R.B. Roy, B. Schwaller, S. P. Walton, J. M. Brandt, D. Tiwari.
“Bringing Differential Privacy to HPC: Privacy-Preserving Transformations of HPC Traces,” In HPDC '25: Proceedings of the 34th International Symposium on High-Performance Parallel and Distributed Computing, 2025.
DOI: hps://doi.org/10.1145/3731545.3731573
Monitoring HPC systems yields valuable insights into user behavior, aiding resource management, collaborative research, and software design. However, privacy concerns raise the barrier for real-world HPC trace sharing between HPC facilities and researchers. Traditional anonymization methods fall short as user behavior remains identifiable. To address this, we propose a robust toolset for privacy protection of HPC traces using Differential Privacy (DP). Our toolset offers a set of DP algorithms, metrics, and visualizations to empower HPC operators to protect users’ sensitive information under a privacy protection guarantee. We evaluated our toolset over real HPC systems traces for different parameters and data aggregations. Moreover, we show that machine learning models trained on privacy-preserved logs maintain accuracy compared to real data, which supports data publishing and sharing across different computing facilities.
S. Viknesh, A. Arzani.
“Differentiable Autoencoding Neural Operator for Interpretable and Integrable Latent Space Modeling,” Subtitled “arXiv:2510.00233v1,” 2025.
Scientific machine learning has enabled the extraction of physical insights from high-dimensional spatiotemporal flow data using linear and nonlinear dimensionality reduction techniques. Despite these advances, achieving interpretability within the latent space remains a challenge. To address this, we propose the DIfferentiable Autoencoding Neural Operator (DIANO), a deterministic autoencoding neural operator framework that constructs physically interpretable latent spaces for both dimensional and geometric reduction, with the provision to enforce differential governing equations directly within the latent space. Built upon neural operators, DIANO compresses high-dimensional input functions into a low-dimensional latent space via spatial coarsening through an encoding neural operator and subsequently reconstructs the original inputs using a decoding neural operator through spatial refinement. We assess DIANO's latent space interpretability and performance in dimensionality reduction against baseline models, including the Convolutional Neural Operator and standard autoencoders. Furthermore, a fully differentiable partial differential equation (PDE) solver is developed and integrated within the latent space, enabling the temporal advancement of both high- and low-fidelity PDEs, thereby embedding physical priors into the latent dynamics. We further investigate various PDE formulations, including the 2D unsteady advection-diffusion and the 3D Pressure-Poisson equation, to examine their influence on shaping the latent flow representations. Benchmark problems considered include flow past a 2D cylinder, flow through a 2D symmetric stenosed artery, and a 3D patient-specific coronary artery. These case studies demonstrate DIANO's capability to solve PDEs within a latent space that facilitates both dimensional and geometrical reduction while allowing latent interpretability.
Y. Wan, H.A. Holman, C. Hansen.
“Beyond flat-panel displays, applications of stereographic and holographic devices in 3D microscopy data analysis,” Subtitled “arXiv:2505.18075,” 2025.
Laser scanning microscopy enables the acquisition of 3D data in biomedical research. A fundamental challenge in visualizing 3D data is that common flat-panel displays, being 2D in nature, cannot faithfully reproduce light fields. Recent years have witnessed the development of various 3D display technologies. These technologies generally fall into two categories, stereography and holography, depending on the number of perspectives they can simultaneously present. We have integrated support for many commercially available 3D-capable displays into FluoRender, a visualization and analysis system for fluorescence microscopy data. This study investigates the opportunities and challenges of applying various 3D display devices in biological research, focusing on their practical use and potential for broad adoption. We found that 3D display devices, including the HoloLens and the Looking Glass, each have their merits and shortcomings. We predict that the convergence of stereographic and holographic technologies will create powerful tools for visualization and analysis in biological applications.
H. Washizaki, N. Aschenbruck, A. Seely, C. Onwubiko, T. Benzel, C. Hansen, E. Au.
“Achievements and Future Directions: IEEE Computer Society 2025 Reflection,” In Computer, IEEE, 2025.
The CS envisions being the global leader in providing technical information, community engagement, and personalized services to computing professionals worldwide. To advance this vision, at the beginning of 2025 the CS reaffirmed its focus on three strategic goals: G1) engaging members, with particular emphasis on students and early career professionals; G2) engaging industry individuals and organizations; and G3) leading in new technical areas.1 These priorities are pursued through programs and activities guided by three operational themes: T1) empowering the volunteer base, T2) enabling nimbleness in execution, and T3) broadening participation. Originally identified by the 2020 CS Planning Committee as the three-year plan, these goals were extended through 2025 to be further reinforced and embedded across the Society.
Page 8 of 152
